


(8 May 2026)
 New Preprint A Regime Theory of Controller Class Selection for LLM Action Decisions at arXiv:2605.06339.
New Preprint A Regime Theory of Controller Class Selection for LLM Action Decisions at arXiv:2605.06339.
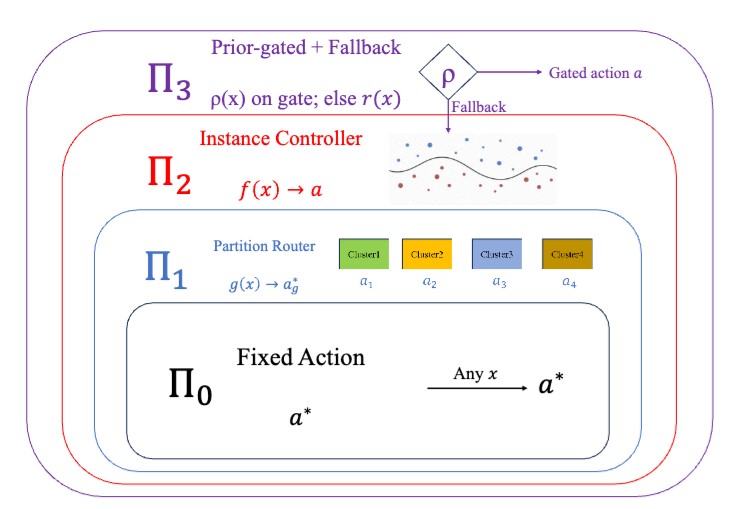
Deployed language and vision-language models must decide, on each input, whether to answer directly, retrieve evidence, defer to a stronger model, or abstain. Contrary to the common monotonicity intuition, greater per-input expressivity is not uniformly beneficial in finite samples - under identical strict cross-validation, different benchmarks prefer different controller classes. This reflects a finite-sample limitation of instance-level uncertainty signals, which can be exhausted at a distribution-dependent scale. We organize controllers into a nested lattice of four classes - fixed actions, partition routers, instance-level controllers, and prior-gated controllers, ordered by complexity. We prove a regime theory that turns three data-estimable bottlenecks into a class choice - how much improvement is possible beyond the best fixed action, whether there are enough samples for instance-level controllers to make reliable decisions, and how much improvement a coarse partition router can recover when instance-level signal is unreliable. The resulting Bernstein-tight threshold has a matching information-theoretic lower bound, and strict nested cross-validation provably selects a near-best class. Across SMS-Spam, HallusionBench, A-OKVQA, and FOLIO, the predicted class matches the empirical winner; the prior-gated controller wins on TextVQA when OCR tokens supply a label-free prediction-time prior.
Read it at arXiv:2605.06339.
(8 May 2026)
 New Preprint BioMedArena - An Open-source Toolkit for Building and Evaluating Biomedical Deep Research Agents at arXiv:2605.06177.
New Preprint BioMedArena - An Open-source Toolkit for Building and Evaluating Biomedical Deep Research Agents at arXiv:2605.06177.
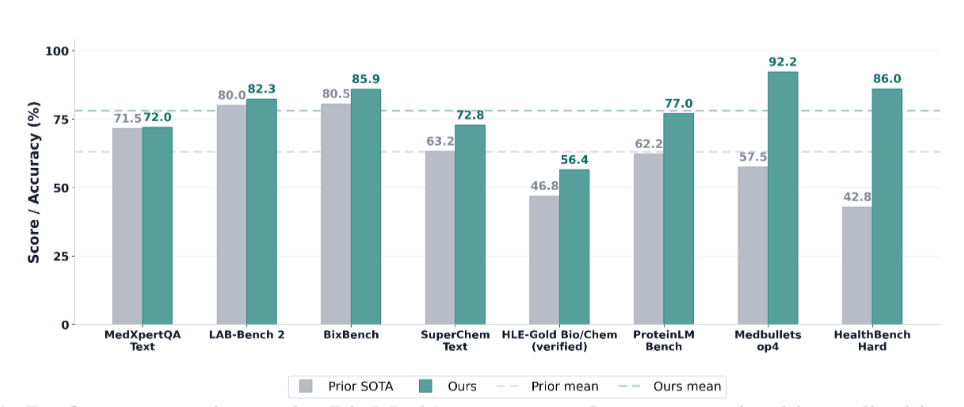
Building a deep research agent today is an exercise in glue code - the same backbone evaluated on the same benchmark can report different accuracies in different papers because harness and tool registry all differ, and integrating a new foundation model into a comparable evaluation surface costs weeks of model-specific engineering. We call this the per-paper engineering tax and release BioMedArena, an open-source toolkit that not only alleviates it but also provides an arena for fair comparison of different foundation models when evaluating them as deep-research agents. BioMedArena decouples six layers of biomedical agent evaluation – benchmark loading, tool exposure, tool selection, execution mode, context management, and scoring – and exposes 147 biomedical benchmarks and 75 biomedical tools across 9 functional families. Adding a new model, benchmark, or tool reduces to registering a few-line provider adapter. We further provide 6 agent harnesses with 6 context-management strategies, which provide 12 backbones with competitive research capabilities and significantly improved performance, achieving state-of-the-art (SOTA) results on 8 representative biomedical benchmarks, with an average lift of +15.03 percentage points over prior SOTA.
Read it at arXiv:2605.06177.
(23 March 2026)
 Paper published on Frontiers in Endocrinology External validation and application of a machine learning–based model for diabetes progression in prediabetes at DOI:10.3389/fendo.2026.1746570.
Paper published on Frontiers in Endocrinology External validation and application of a machine learning–based model for diabetes progression in prediabetes at DOI:10.3389/fendo.2026.1746570.
This study externally validated a machine learning–based model for type 2 diabetes progression (ML-PR) and evaluated its clinical utility in individuals with prediabetes. We included 3,081 participants from the Diabetes Prevention Program (DPP) and the DPP Outcome Study (DPPOS). The ML-PR model was assessed using dicrimination, calibration curves, and decision curve analysis, and its performance was compared with existing diabetes prediction models. Based on ML-PR scores, patients were stratified into high- or low-risk categories. Cox proportional hazards and logistic regression models were used to evaluate the incidence of type 2 diabetes, microvascular complications, and cardiovascular events across risk and intervention groups. The ML-PR model achieved an area under the ROC curve of 0.74 (95% confidence interval 0.71–0.78) for predicting 3-year progression to type 2 diabetes. Calibration and decision curve analyses indicated good agreement and net clinical benefit. High-risk individuals exhibited a significantly higher risk of developing type 2 diabetes in both the DPP and DPPOS cohorts (P < 0.001), as well as a 67% increased risk of microvascular complications in DPPOS (P < 0.001), though no significant difference in cardiovascular risk was observed. Significant interactions between treatment and risk group were identified, indicating that high-risk participants benefited more from lifestyle modification and metformin interventions (P for interaction = 0.03 in DPP; P = 0.014 in DPPOS).
Read it at DOI:10.3389/fendo.2026.1746570.
(12 March 2026)
 New Preprint LoV3D - Grounding Cognitive Prognosis Reasoning in Longitudinal 3D Brain MRI via Regional Volume Assessments at arXiv:2603.12071.
New Preprint LoV3D - Grounding Cognitive Prognosis Reasoning in Longitudinal 3D Brain MRI via Regional Volume Assessments at arXiv:2603.12071.
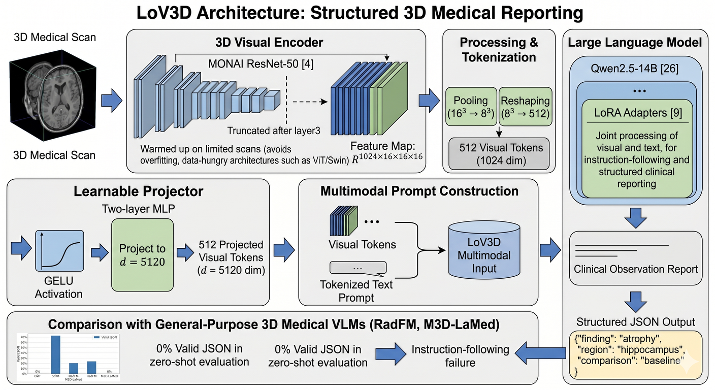
Longitudinal brain MRI is essential for characterizing the progression of neurological diseases such as Alzheimer’s disease assessment. However, current deep-learning tools fragment this process - classifiers reduce a scan to a label, volumetric pipelines produce uninterpreted measurements, and vision-language models (VLMs) may generate fluent but potentially hallucinated conclusions. We present LoV3D, a pipeline for training 3D vision-language models, which reads longitudinal T1-weighted brain MRI, produces a region-level anatomical assessment, conducts longitudinal comparison with the prior scan, and finally outputs a three-class diagnosis (Cognitively Normal, Mild Cognitive Impairment, or Dementia) along with a synthesized diagnostic summary. The stepped pipeline grounds the final diagnosis by enforcing label consistency, longitudinal coherence, and biological plausibility, thereby reducing the risks of hallucinations. The training process introduces a clinically-weighted Verifier that scores candidate outputs automatically against normative references derived from standardized volume metrics, driving Direct Preference Optimization without a single human annotation. On a subject-level held-out ADNI test set (479 scans, 258 subjects), LoV3D achieves 93.7% three-class diagnostic accuracy (+34.8% over the no-grounding baseline), 97.2% on two-class diagnosis accuracy (+4% over the SOTA) and 82.6% region-level anatomical classification accuracy (+33.1% over VLM baselines). Zero-shot transfer yields 95.4% on MIRIAD (100% Dementia recall) and 82.9% three-class accuracy on AIBL, confirming high generalizability across sites, scanners, and populations.
Read it at arXiv:2603.12071.
(12 March 2026)
 New Paper Natural language processing for geriatric syndromes - a systematic review of methods, applications, and challenges published at DOI:10.1186/s12911-026-03417-0.
New Paper Natural language processing for geriatric syndromes - a systematic review of methods, applications, and challenges published at DOI:10.1186/s12911-026-03417-0.
Geriatric syndromes (GS) are complex conditions that affect older adults and often require multidisciplinary assessment. Natural language processing (NLP) has emerged as a promising tool for extracting relevant clinical information from unstructured text in electronic health records (EHRs). However, the application of NLP in detecting and monitoring GS remains an evolving area of research. This systematic review explores the role of NLP in the identification and analysis of GS, examining its applications, methodologies, and effectiveness. Furthermore, this review discusses the existing challenges, limitations, and future directions to advance NLP applications in the GS research.
Read it at DOI:10.1186/s12911-026-03417-0.
(25 February 2026)
 Paper published on Lancet Digital Health RareArena - a comprehensive benchmark dataset unveiling the potential of large language models in rare disease diagnosis at 10.1016/j.landig.2025.100953
Paper published on Lancet Digital Health RareArena - a comprehensive benchmark dataset unveiling the potential of large language models in rare disease diagnosis at 10.1016/j.landig.2025.100953
The study introduces RareArena, a new large benchmark dataset designed to improve how artificial intelligence (AI), especially large language models (LLMs), is evaluated for rare disease diagnosis. Medical AI has shown promise in interpreting clinical text and supporting diagnosis, but existing evaluations often rely on small or artificial test sets that don’t reflect the real complexity doctors face. RareArena tackles this gap by collecting thousands of real clinical case reports from PubMed Central and organising them into tasks that mimic real-world clinical problem-solving — such as screening for rare diseases and confirming specific diagnoses from free-text descriptions. The dataset enables systematic comparison of AI models’ abilities to understand medical language and recognise rare conditions, highlighting strengths and limitations in current systems. By providing a realistic testing ground, RareArena aims to accelerate the development of safer, more trustworthy AI tools that can support clinicians in identifying rare diseases earlier and more accurately.
Read it at 10.1016/j.landig.2025.100953.
(3 February 2026)
 Paper published on Radiology Guidelines for Reporting Studies on Large Language Models in Radiology - An International Delphi Expert Survey at https://pubs.rsna.org/doi/full/10.1148/radiol.250913
Paper published on Radiology Guidelines for Reporting Studies on Large Language Models in Radiology - An International Delphi Expert Survey at https://pubs.rsna.org/doi/full/10.1148/radiol.250913
Large language models (LLMs) have transformative potential in radiology, including textual summaries, diagnostic decision support, proofreading, and image analysis. However, the rapid increase in studies investigating these models, along with the lack of standardized LLM-specific reporting practices, affects reproducibility, reliability, and clinical applicability. To address this, reporting guidelines for LLM studies in radiology were developed using a two-step process. First, a systematic review of LLM studies in radiology was conducted across PubMed, IEEE Xplore, and the ACM Digital Library, covering publications between May 2023 and March 2024. Of 511 screened studies, 57 were included to identify relevant aspects for the guidelines. Then, in a Delphi process, 20 international experts developed the final list of items for inclusion. Items consented as relevant were summarized into a structured checklist containing 32 items across six key categories - general information and data input; prompting and fine-tuning; performance metrics; ethics and data transparency; implementation, risks, and limitations; and further/optional aspects. The final FLAIR (Framework for LLM Assessment in Radiology) checklist aims to standardize reporting of LLM studies in radiology, fostering transparency, reproducibility, comparability, and clinical applicability to enhance clinical translation and patient care..
Read it at https://pubs.rsna.org/doi/full/10.1148/radiol.250913.
(16 January 2026)
 Preprint Prevalence of 406 rare diseases by ethnicity and their associated COVID-19 infection burden - A national cross-sectional study of 62.5 million people in England at https://www.medrxiv.org/content/10.64898/2026.01.13.26344068v1
Preprint Prevalence of 406 rare diseases by ethnicity and their associated COVID-19 infection burden - A national cross-sectional study of 62.5 million people in England at https://www.medrxiv.org/content/10.64898/2026.01.13.26344068v1
The burden of the COVID-19 pandemic disproportionately affected individuals with rare diseases. However, the patterning of this risk by ethnicity is complex and runs contrary to general population trends, likely reflecting the deep-seated ethnic disparities in the prevalence of specific RDs. Our foundational map of 406 rare diseases by granular ethnicity is essential for understanding these factors and identifying which specific patient-ethnic subgroups face the greatest intersectional risk.
Read the preprint version at https://www.medrxiv.org/content/10.64898/2026.01.13.26344068v1.
(4 November 2025)
 Paper accepted by AAAI 2026 Artificial Intelligence for Social Impact Track – Error Correction in Radiology Reports - A Knowledge Distillation-Based Multi-Stage Framework at https://arxiv.org/abs/2406.15045v3
Paper accepted by AAAI 2026 Artificial Intelligence for Social Impact Track – Error Correction in Radiology Reports - A Knowledge Distillation-Based Multi-Stage Framework at https://arxiv.org/abs/2406.15045v3
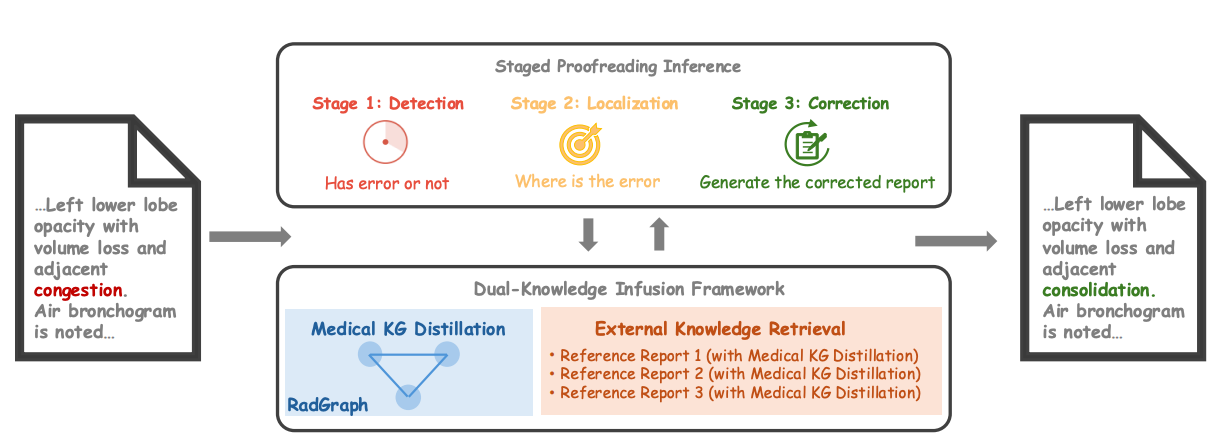
The increasing complexity and workload of clinical radiology leads to inevitable oversights and mistakes in their use as diagnostic tools, causing delayed treatments and sometimes life-threatening harms to patients. While large language models (LLMs) have shown remarkable progress in many tasks, their utilities in detecting and correcting errors in radiology reporting are limited. We present a novel dual-knowledge infusion framework that enhances LLMs’ capability for radiology report proofreading through systematic integration of medical expertise. Specifically, our knowledge infusion combines medical knowledge graph distillation (MKGD) with external knowledge retrieval (EXKR), enabling an effective automated approach in tackling mistakes in radiology reporting. By decomposing the complex proofreading task into three specialized stages of detection, localization, and correction, our method mirrors the systematic review process employed by expert radiologists, ensuring both precision and clinical interpretability. The dual-knowledge framework captures intricate medical relationships through structured graph representations while leveraging curated clinical expertise from reference reports. To perform a robust, clinically relevant evaluation, we constructed a comprehensive benchmark using real-world radiology reports with error patterns derived from real-world scenarios, including speech recognition confusions, terminology ambiguities, and template-related inconsistencies, all validated by practicing radiologists. Extensive evaluations across multiple LLM architectures demonstrate substantial improvements of our approach - up to 31.56% increase in error detection accuracy and 37.4% reduction in processing time. Human evaluation by radiologists confirms superior clinical relevance and factual consistency compared to existing approaches. Our framework addresses critical needs in clinical practice by enhancing report quality while reducing radiologist burden, particularly benefiting resource-constrained healthcare environments.
Read the preprint version at https://arxiv.org/abs/2406.15045v3.
(31 October 2025)
 A systematic review published on Communications Medicine - Digital health technology use in clinical trials of rare diseases - a systematic review at https://www.nature.com/articles/s43856-025-01137-6
A systematic review published on Communications Medicine - Digital health technology use in clinical trials of rare diseases - a systematic review at https://www.nature.com/articles/s43856-025-01137-6
Hundreds of millions of people around the world live with rare diseases, but most of these conditions still don’t have effective treatments. Clinical trials are vital for new therapies, but rare disease trials are especially challenging due to small, dispersed patient populations and difficulties with long-term participation. This study looked at how digital health tools—like wearable devices, mobile apps, and healthcare platforms—are being used to improve rare disease clinical trials. We analysed 262 trials across ten rare diseases and found that digital health technologies were increasingly used for remote monitoring, digital treatment, and long-term care. By making it easier for patients to participate in trials—no matter where they live—digital technologies can help speed up the development of much-needed treatments and help make research more inclusive and equitable by reaching patients who would otherwise be left out.
Read the full article at https://www.nature.com/articles/s43856-025-01137-6.
(21 October 2025)
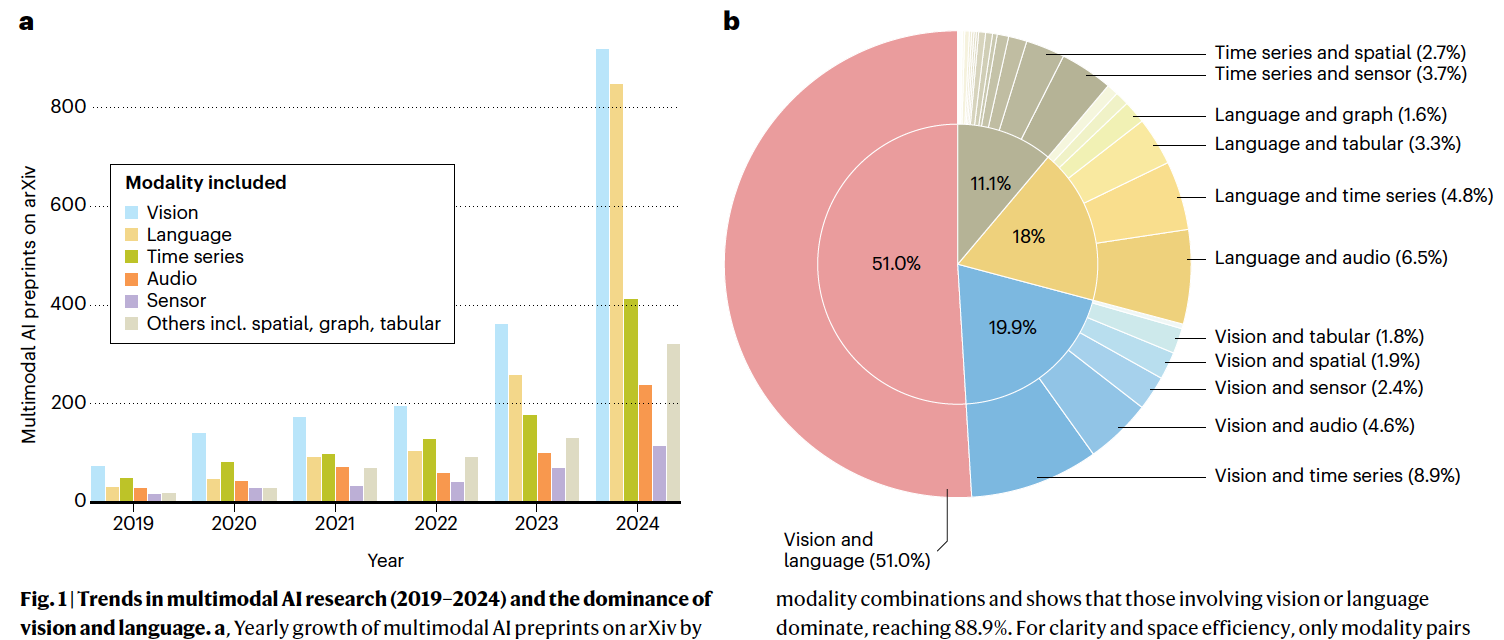 A collaborative, perspective paper published on Nature Machine Intelligence - Towards deployment-centric multimodal AI beyond vision and languag at https://www.nature.com/articles/s42256-025-01116-5
A collaborative, perspective paper published on Nature Machine Intelligence - Towards deployment-centric multimodal AI beyond vision and languag at https://www.nature.com/articles/s42256-025-01116-5
Multimodal artificial intelligence (AI) integrates diverse types of data via machine learning to improve understanding, prediction and decision-making across disciplines such as healthcare, science and engineering. However, most multimodal AI advances focus on models for vision and language data, and their deployability remains a key challenge. We advocate a deployment-centric workflow that incorporates deployment constraints early on to reduce the likelihood of undeployable solutions, complementing data-centric and model-centric approaches. We also emphasize deeper integration across multiple levels of multimodality through stakeholder engagement and interdisciplinary collaboration to broaden the research scope beyond vision and language. To facilitate this approach, we identify common multimodal-AI-specific challenges shared across disciplines and examine three real-world use cases - pandemic response, self-driving car design and climate change adaptation, drawing expertise from healthcare, social science, engineering, science, sustainability and finance. By fostering interdisciplinary dialogue and open research practices, our community can accelerate deployment-centric development for broad societal impact.
Read the full article at https://www.nature.com/articles/s42256-025-01116-5.
(19 September 2025)
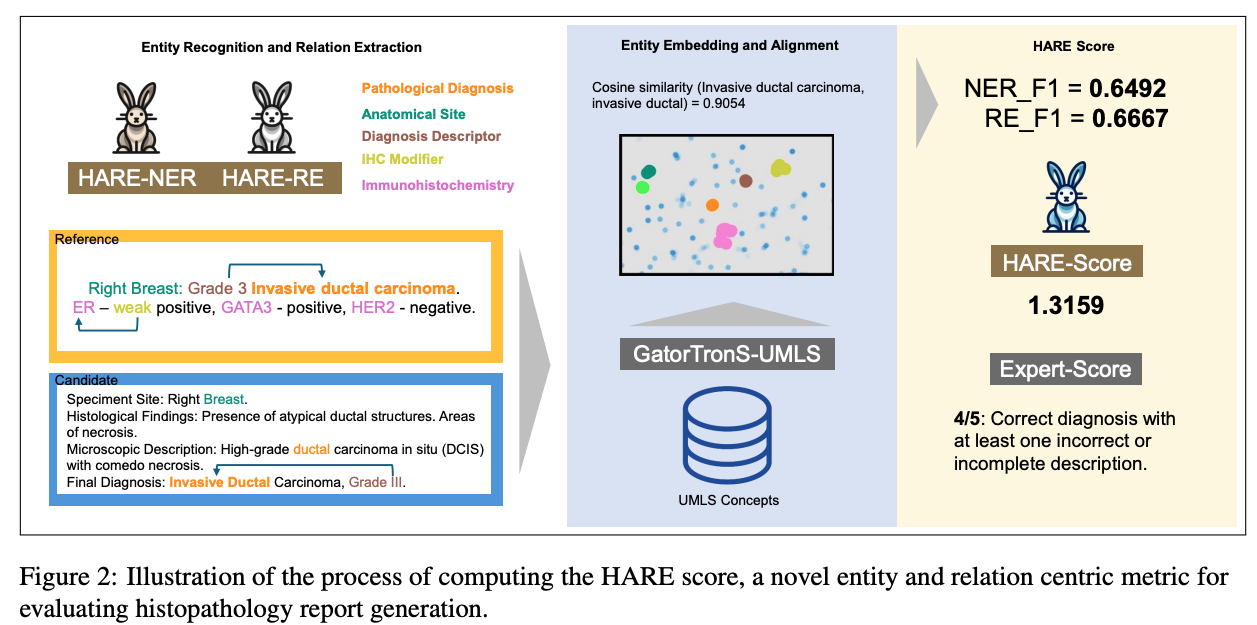 Paper accepted by EMNLP 2025 findings – HARE- an entity and relation centric evaluation framework for histopathology reports – at https://arxiv.org/abs/2507.09097
Paper accepted by EMNLP 2025 findings – HARE- an entity and relation centric evaluation framework for histopathology reports – at https://arxiv.org/abs/2507.09097
Large Vision-Language Models (LVLMs) have demonstrated promising performance in chest X-ray (CXR) analysis. To enhance human-computer interaction, several studies have incorporated radiologists’ eye gaze, typically through heatmaps or textual prompts. However, these methods often overlook the sequential order of eye movements, which could provide valuable insights by highlighting both the areas of interest and the order in which they are examined. In this work, we propose a novel approach called RadEyeVideo that integrates radiologists’ eye-fixation data as a video sequence, capturing both the temporal and spatial dynamics of their gaze. We evaluate this method in CXR report generation and disease diagnosis using three general-domain, open-source LVLMs with video input capabilities. When prompted with eye-gaze videos, model performance improves by up to 24.6% in the report generation task and on average 15.2% for both tasks using scaled evaluation metrics. Notably, RadEyeVideo enhanced an open-domain LVLM model, LLaVA-OneVision, to surpass task-specific medical LVLMs such as MAIRA-2 and CheXagent, trained on large Chest X-ray data. This work highlights that domain expert’s knowledge (eye-gaze information in this case), when effectively integrated with LVLMs, can significantly enhance general-domain models’ capabilities in clinical tasks. RadEyeVideo is a step toward a scalable human-centered approach of utilizing LVLMs in medical image analytics.
Read the full paper at https://arxiv.org/abs/2507.09097.